Telomeres are repetitive nucleotide sequences at the end of the chromosomes that protect chromosome ends from degradation and fusion. Telomeres continuously lose their sequence repeats during cellular replication, so this progressive telomere shortening becomes a biomarker for ageing as a molecular clock. In this article, the authors review the recent progress on the structure, the measurement technology, the physiology, and the pathology of telomeres. The authors also discuss the activation of telomerase in the context of rejuvenation and control of age-related diseases in human and animal models. The association of telomeres and stem cells is also discussed.
A telomere is the region of repetitive nucleotide sequences at each end of a chromatid, which protects the ends of the chromosome from deterioration or from fusion with neighbouring chromosomes1. Its name is derived from the Greek nouns telos ‘end’ and meros ‘part’. The nucleotide sequence varies in different phyla, but in human telomeres, the repetitive unit is TTAGGG and spans around 5–15 kilobases (Kb).
Telomeres continuously lose these repeat sequences during cellular replication, owing to the incomplete replication of linear chromosomes by conventional DNA polymerases; a feature known as ‘the end‑replication problem’2. This progressive telomere shortening confers the idea of a ‘molecular clock’, which thus provides a useful biomarker for ageing research. Telomere shortening to a critical length causes chromosomal dysfunctions such as chromosomal instability, chromosomal fusion or chromosomal deletion, and results in loss of cell viability called cellular senescence3. Dozens of proteins have been reported to associate with telomeres, including telomere-binding proteins, DNA repair proteins, and telomerase that specifically recognises 3´ OH groups at the end of the telomere structure4. Telomerase is a reverse transcriptase that synthesises new telomere sequences onto chromosome ends. This enzyme is specifically expressed in germ line cells, somatic stem cells and cancer cells, but not in the majority of somatic cells1. Telomeres are also characterised by 150–200 nucleotide single-strand overhang of the G-rich strand that folds back and anneals with the double-stranded region of the TTAGGG repeats to form a large telomeric loop known as the T-loop and a short loop known as the D-loop. This complex structure composed of TTAGGG repeats, binding proteins, T-loop, D-loop, and telomerase contributes to the functional integrity and the protection of chromosome ends. Supporting this idea, the loss of TTAGGG repeats or telomere-binding proteins all result in chromosomal dysfunction, such as chromosomal fusions4.
Telomere length measurement
Traditionally, telomere length measurement has relied on terminal restriction fragment (TRF), which is the application of the classical southern-blot technique to detect telomere sequences. This approach provides a visual representation of the distribution of telomere lengths in the cell population. Although considered to be a robust test, its sensitivity is relatively poor and results depend on the total DNA amount and integrity, and electrophoretic conditions5.
More recently, a new strategy based on quantitative polymerase chain reaction (Q-PCR) has replaced TRF for clinical studies6. The advantages of the Q-PCR method are that it is relatively fast, requires minute amounts of DNA, and is capable of high throughput. The disadvantage is that it does not provide information about individual cells, so the results represent averages of telomere length. In addition, sample quality heavily compromises the relative measurements of telomere sequence to that of single-copy genes used for normalisation, introducing significant errors in telomere length measurement.
With the advent of peptide nuclei acid probes, the fluorescence in situ hybridisation (FISH) technique evolved into a quantitative tool to measure individual telomeres in nuclei. Yet, the standard Q-FISH technique only allows the detection and quantification of telomeres in condensed chromosomes of mitotic cells. The need for cells in mitosis limits the method’s scalability to the required level for epidemiological and clinical studies.
In 2007, Canela et al7 developed a high-throughput (HT) Q-FISH test that overcame the limitations of the classical assay. In this study, it was shown that HT Q-FISH could be applied on interphase cells giving similar results to those of a standard Q-FISH. Both standard and HT Q-FISH measurements perfectly correlated with those of TRF, attesting the robustness of the new assay.
HT Q-FISH was licensed to the Spanish company Life Length for further development into a commercial tool. This method allows visualisation and quantitation in a HT confocal microscope of individual telomeres within the cells, allowing the determination of the entire telomere length distribution of the sample. Knowing the distribution of telomere lengths in the cells’ chromosomes is of upmost importance for two reasons: it gives insight into the fraction of the shortest telomeres, which are regarded as the principal cause of replicative cell ageing and age-related diseases8; and it permits the calculation of the median and modal telomere length distribution, which are more representative of the telomere length in cells than the average, as the distribution of telomere lengths is not normal but skewed (Figure 1B).
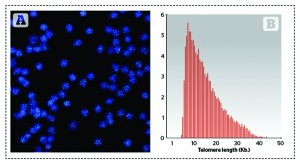
Figure 1. Q-FISH image and frequency distribution of telomere length in cells. (A) Image of Q-FISH taken on purified lymphocytes. Nuclei were stained with DAPI (blue) and telomeres are visualised in green. (B) Example of the distribution of telomere lengths in human lymphocytes
This advantage of HT Q-FISH has been illustrated in an integrative personal omics profile studied on an individual who contained a mutation in his TERT gene. Despite this mutation, the subject presented a median telomere length undistinguishable from TERT+ individuals. However, he also presented an increased percentage of short telomeres compared to the normal population9. Owing to the scalability of the technique, as well as its high sensitivity and the wealth of information provided, HT Q-FISH is emerging as the method of choice to carry out more sensitive studies on telomere length.




